IntroductionI haven't posted in a while because I have been trying to find new and exciting papers a little out of my comfort zone. And it just so happened that Scripps hosted National Academy member, Diane Newman, this week for a special seminar sponsored by the graduate students in the Biotechnology Training Fellowship. Almost all of my interest has been driven by a graduate student in my current lab, Doug Sweeney, who is actively looking at the mechanisms of electron transfer in marine sediment bacterium. If you are interested in any of these topics, definitely feel free to email him!
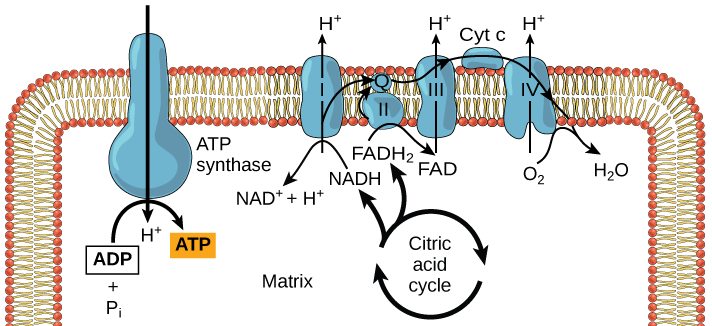
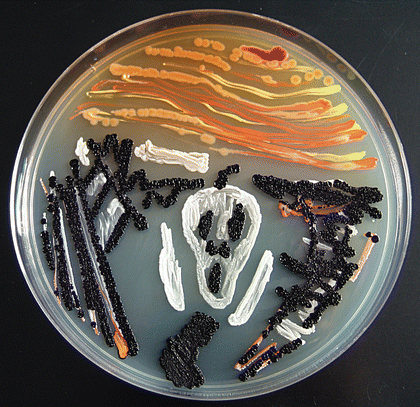
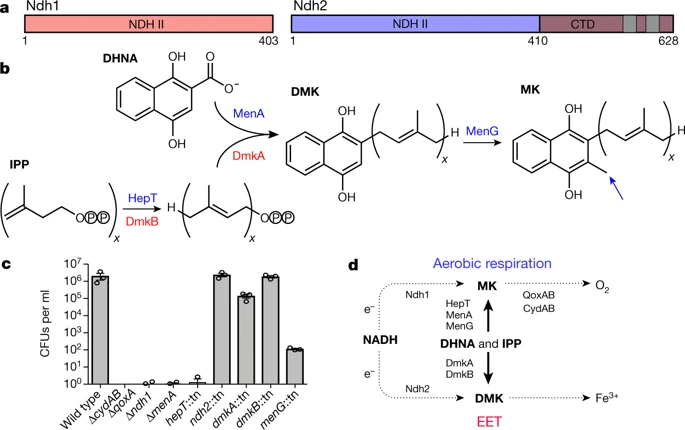
Redox potential are everything in microbial communitiesBefore I jump into the chemistry side of things, I first wanted to give a brief overview of why electron transfer is so imperative. Most of you probably know this, but this process of transferring electrons enables cells to maintain cellular function and generate energy for additional processes. But since I am not a chemist, I obviously want to take a step back very briefly to highlight how important these processes can be at multiple scales, especially when we consider entire communities! A recent paper by Ramirez-Flandes et al. (2019) assessed metagenomes from 18 biomes to determine what distinguishing characteristics delineate microbial communities [1]. Most of us are aware that taxonomic signatures can illuminate certain microbial biomes (e.g., Cyanobacteria are usually associated with marine samples). However, this paper decided to turn to functional characterization to differentiate various microbial biomes. What they found was that oxidoreductase gene profiles separated the biomes into three main clusters: anoxic biomes (e.g., host-associated and marine sediments), aquatic biomes, and soil-associated biomes. Within the oxidoreductase gene profiles, different biomes were associated with various groups of oxidoreductases related to substrate availability and stress conditions. Redox processes alone delineate microbial communities! Why are redox processes so important?I started with this first paper because it nicely outline that various biomes present various challenges for the bacteria that reside there. For example, in the absence of oxygen, how do organisms maintain their cellular processes? One way is fermentation, but this leads to toxic byproducts and is not a sustainable option. Another is the use of alternative electron acceptors. In soils, sediments, host-associated, microbes are constantly dealing with anoxic conditions and yet we find traditionally described aerobic organisms. Some organisms have adapted to utilize redox-active minerals that are abundant in certain biomes, like iron or manganese in soils. This is absolutely fascinating, microbes have evolved novel strategies to exchange electrons with minerals by transferring electrons from the cytosol to the exterior of the cell [for more see this review [2]]. So how do these microbes transfer electrons outside the cell? The above image [3] depicts the beautiful pigments bacteria are capable of producing on a plate that most of us take for granted. But what is the role of these pigments? Some may be related to UV or desiccation, but others often change colors which is indicative of a redox reaction. This led to the topic of the entire post, the use of these molecules as extracellular electron shuttles. Some known shuttles are the natural products, quinones, phenazines, and flavins, which shuttle electrons to an acceptor. Glasser et al. [3] found that phenazines in particular are widespread along with the transcription factor, SoxR, that senses redox-active metabolites. From Dr. Newman's seminar this week, she discussed (in fascinating detail) how these phenazines in biofilms interact with eDNA (or dead cell DNA) to stabilize redox reactions and allow for the anaerobic survival of the producing organisms to maintain cellular viability. These discoveries really revolutionize our understanding of microbial metabolism. But the biggest question I always had is the production of such natural products must be expensive, especially considering their role is essential in stressed conditions. And even if production is sustained, how can the cell ensure an adequate return on its investment; in other words, how can the extracellular molecule not be lost to the environment. Dr. Newman presented an amazing solution to this problem by addressing shuttle diffusion gradients, electron hopping, and more [see her review [3] and stay tuned for some forthcoming work from her lab]. Why do microbes go through all this trouble???Finally, this question brings me to the Journal Club paper of the week [4]. Why are microbes evolving these intricate solutions to a seemingly simple problem of where to deposit electrons. As a reminder, the entire point of transferring electrons is to generate ATP, but to do so requires the use of electron shuttles such as NADH to carry electrons from one reaction to another. This essential molecule then can be used as a reducing agent to donate electrons and become oxidized as NAD+, ready to accept more electrons and continue the cycle. In the absence of a terminal electron acceptor, such as oxygen, NADH will build up in the cell. What would be a good strategy to deal with NADH buildup? Well in a paper by Light et al. [4] the authors identified a parallel electron transfer pathway in anaerobic conditions in the pathogen Listeria monocytogenes. In the presence of oxygen, normal aerobic respiration occurs with an essential enzyme for metabolism, NADH dehydrogenase (Ndh1 in above figure), catalyzing electron exchange to a quinone derivative, the first step in the electron transport chain. Under anaerobic conditions, this process cannot continue as NADH cannot be recycled. So, a parallel solution evolved using another NADH dehydrogenase (Ndh2 in above figure) enables NAD+ concentrations to be restored. Using transposon mutants the authors show cell viability using knockouts of these essential genes (panel C); for instance, in the presence of oxygen the knockout of Ndh2 (or any other genes in the anaerobic pathway) does not inhibit cellular growth. Under anaerobic conditions, the demethylated quinone derivative (DMK) accepts electrons from NADH via Ndh2. The authors very thoroughly outline the subsequent electron transferring system where DMK donates electrons to membrane proteins and finally to environmental flavins to shuttle electrons and finally to extracellular acceptors, like Fe(III). More and more, it appears that these innovative strategies to deal with electron transfer are a fundamental issue in all microbial life, not limited to specialized processes in mineral-respiring bacteria. All bacteria require cell viability and processes. These processes are driven entirely by redox potential which is mediated by electron shuttles. This really blows my mind as I have never considered the magnitude of the electron transport chain (and the various innovations) to deal with the recycling of electron shuttles. Papers:1. Ramirez-Flandes S, Gonzalez B, Ulloa O. (2019). Redox traits characterize the organization of global microbial communities. Proceedings to the National Academy of Sciences 116(9): 3630-3635.
2. Shi L, Dong H, Reguera G, Beyenal H, Lu A, Liu J, Yu HQ, Frederickson JK. (2016). Extracellular electron transfer mechanisms between microorganisms and minerals. Nature Reviews Microbiology 14: 651-662. 3. Glasser NR, Saunders SH, Newman DK. (2017). The colorful world of extracellular electron shuttles. Annual Review of Microbiology 71: 731-751. 4. Light SH, Su L, Rivera-Lugo R, Cornejo JA, Louie A, Iaverone AT, Ajo-Franklin CM, Portnoy DA. (2018). A flavin-based extracellular electron transfer mechanism in diverse Gram-positive bacteria. Nature 562: 140-144.
0 Comments
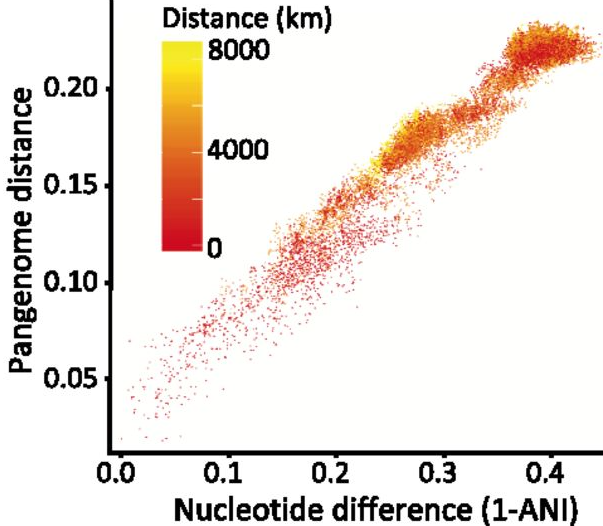
Introduction to microbial biogeographyUtilizing a biogeographic framework is extremely powerful as biologists can understand the ecological and environmental factors driving the distribution of taxa. In addition, extensions of this include phylogeography where evolutionary biologists can understand barriers to dispersal and geological events that may contribute to species' distributions. More recently, these patterns have been observed for microbes. However, microbial biogeographic patterns are typically associated with environmental factors at the community level. This is in stark contrast to the traditional work in plants and animals, where species' distributions and/or populations are described. Previously, I wrote a commentary with my PhD advisor, Jennifer Martiny, going over some of these ideas [1]. Recently, I have seen a lot more papers moving beyond community-level patterns and trying to identify the ecological and evolutionary processes structuring the distribution of microbial taxa. So, for this month, I thought everyone could take a deep dive into some of these amazing papers. I tried to find papers referencing soil, marine, and host-associated bacteria to give a little bit of something for everyone. This list is in no way inclusive, so please feel free to share other papers you have seen as well! I will be more than happy to add to this list. Global-level patternsFirst couple of papers I want to highlight are looking at the global distribution of microbial taxa. To start, we can look at soil bacteria. A lot of papers have previously shown that, at global levels, various edaphic properties can structure soil communities, including pH, temperature, and salinity. Greenlon et al. examined the phylogenetic diversity of the nodule-forming genus, Mesorhizobium, by sampling chickpeas across various soil types, climates, continents, etc. [2].
The second paper dives into maybe the most abundant organism on the planet, the SAR11 clade. This order has a vast global distribution in oceans with finer clades exhibiting differential geographic patterns. Despite its ubiquity and global distribution, SAR11 remains incredibly difficult to culture and also creates problems reconstructing MAGs from environmental surveys. As such, this next paper attempts to investigate the genomic diversity of the SAR11 clade using read recruitment of oceanic samples to identify single-amino acid variants [3]. One issue I keep coming across in any investigation into the SAR11 clade is that most studies are examining genetic diversity across an entire order. Is it really surprising that different families, genera, species, populations (if we can get to this level) within the SAR11 order have differential distributions? And why is it applicable to then look at non-synonymous variation? In any case, the authors further extrapolate amino acid variants to physico-chemical properties to conclude that the hydrophobic interactions of amino acids are under purifying selection. Another question I had how many amino acid positions would be affected by the chemistry, is this restricted to the active sites? How did they tease a part neutral divergence in these proteins across divergent taxa? Finally, the paper concludes that proteotypes within a SAR11 subclade exhibit differential biogeographic patterns with temperature being a driving determinant of spatial partitioning. But then they conclude that "lending credence to the idea for marine systems that 'everything is everywhere but the environment selects'"? - shouldn't these distinct biogeographic patterns reflect the opposite? Regional biogeographic patternsRates of homologous recombination between closely-related strains have largely been inferred from genome sequences, but Potnis et al. utilize in vitro recombinant strains along with wild-type isolates to determine the extent of HR [4]. The authors identified large recombining fragments (upwards of 30kbp) in concentrated genomic islands. Functional annotations of these regions revealed genes related to the transition of pathogenic lifestyles of the plant pathogen.
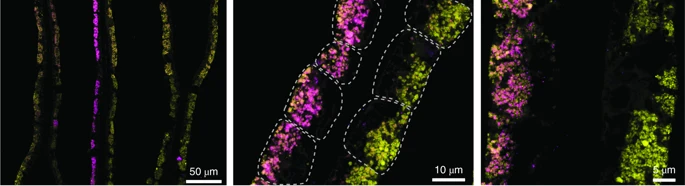
Fine spatial scaleStaying with soil dynamics, next I want to look at this really neat paper that sampled Streptomyces strains at spatial microscales (mm to cm). Similarly to the SAR11 clade, this "genus" encompasses extensive genetic diversity. Based on these ambiguous delineations, I think a lot of studies have overestimated the extent of HGT in structuring Streptomyces diversity; actually a recent study quantified this showing they are indeed rare HGT events across distantly-related Strepto lineages. Back to this paper which isolated 32 strains from soil aggregates that shared >98.6% ANI [6]. Despite this, conspecific strains harbored extensive gene content differences in the flexible genome and proposed that the massive gene fluxes are probably mediated by Actino ICEs (AICEs). Some of these indels included biosynthetic gene clusters (BGCs) - this might only be a cool result for me! In conclusion, the observed gene fluxes between microscale spatially-separated strains was mediated by frequent transfer among closely-related strains via conjugation. This last paper is really interesting as **SPOILER** they show as many as 16?!?!? strains of intracellular symbionts can coexist in a single individual of deep sea mussels [7]. I find this result fascinating since we would expect highly similar strains to outcompete one another via the competitive exclusion principle. In this system, it was thought the mussels hosted two intracellular symbionts, one for sulfur-oxidizing (SOX) and another for methane-oxidizing - both contribute energy sources for carbon fixation. By 16S rRNA standards, each mussel host was thought to harbor a single symbiont species of SOX and methane oxidizers. This paper utilized metagenomes and metatranscriptomes to identify variable strain diversity in a single mussel host. Further, these variations were linked to strain-specific genes related to key functional processes in the SOX symbionts. For instance, some symbiont strains expressed genes related to H2 oxidation; the authors also confirmed with imaging analysis (below). This imaging revealed such fine-scale spatial partitioning that it was restricted to the host bacteriocytes - that is incredible compartmentalization! Simultaneous FISH of hydrogenase operon (violet) and 16S rRNA (green) of the SOX symbiont in gill tissue of B. azoricus from site encoding genes related to H2 oxidation. Papers:1. Chase AB, Martiny JBH. (2018). The importance of resolving biogeographic patterns of microbial microdiversity. Microbiology Australia 39(1): 5-8.
2. Greenlon A, Chang PL, Damtew ZM, Muleta A, Carrasquilla-Garcia N, Kim D, Nguyen HP, Suryawanshi V, Krieg CP, Yadav SM, Patel JS, Mukherjee A, Udupa S, Benjelloun, Thami-Alami I, Yasin M, Patil B, Singh S, Sarma BK, von Wettberg EJB, Kahraman A, Bukun B, Assefa F, Tesfaye K, Fikre A, Cook DR. (2019). Global-level population genomics reveals differential effects of geography and phylogeny on horizontal gene transfer in soil bacteria. Proceedings to the National Academy of Sciences 116(30): 15200-15209. 3. Delmont TO, Kiefl E, Kilinc O, Esen OC, Uysal I, Rappe MS, Giovannoni S, Eren AM. (2019). Single-amino acid variants reveal evolutionary processes that shape the biogeography of a global SAR11 subclade. eLife 8:e46497. 4. Potnis N, Kandel PP, Merfa MV, Retchless AC, Parker JK, Stenger DC, Almeida RPP, Bergsma-Vlami M, Westenberg M, Cobine PA, de la Fuente L. (2019). Patterns of inter- and intrasubspecific homologous recombination inform eco-evolutionary dynamics of Xylella fastidosa. The ISME Journal 13: 2319–2333. 5. Chase AB, Arevalo P, Brodie EL, Polz MF, Karaoz U, Martiny JBH. (2019). Maintenance of sympatric and allopatric populations in free-living terrestrial bacteria. mBio 10(5): e02361-19. 6. Tidjani A, Lorenzi J, Toussaint M, van Dijk E, Naquin D, Lespinet O, Bontemps C, Leblond P. (2019). Massive gene flux drives genomic diversity between sympatric Streptomyces conspecifics. mBio 10(5): e01533-19. 7. Ansorge R, Romano S, Sayavedra L, Porras MAG, Kupczok A, Tegetmeyer HE, Dubilier N, Peterson J. (2019). Functional diversity enables multiple symbiont strains to coexist in deep-sea mussels. Nature Microbiology |
AuthorSome thoughts on some (small) things Archives
May 2023
Categories |
Proudly powered by Weebly

 RSS Feed
RSS Feed