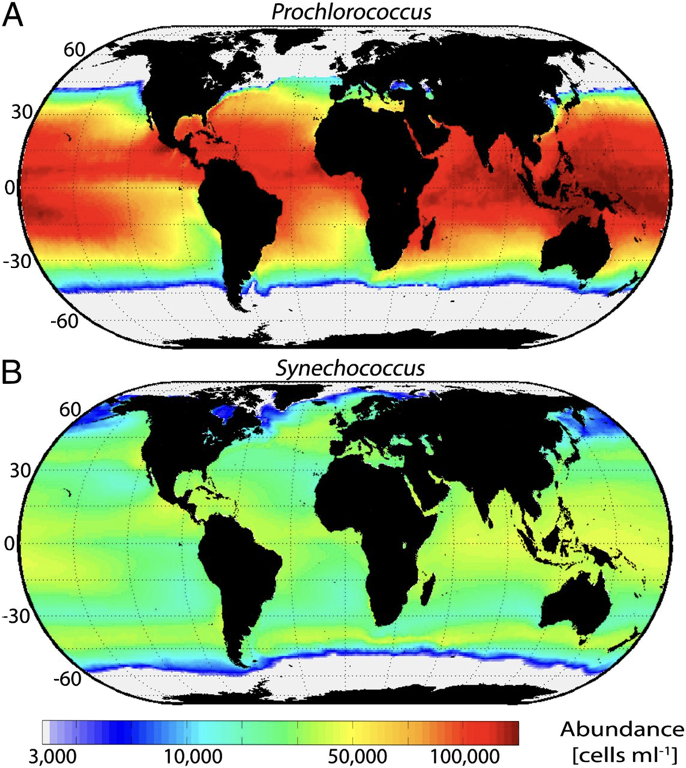
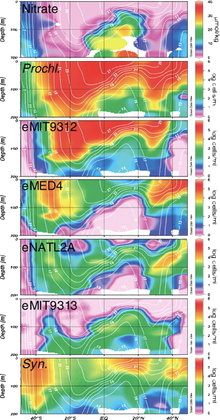
BackgroundWhy are there so many species and what are the limitations for the coexistence of N species? These are fundamental questions in ecology and evolution that have been at the foundation of these fields for centuries. On one hand, Hubbell's Neutral Theory [1] assumes that all species are essentially ecologically equivalent (does not actually claim this, just that neutral processes are greater than deterministic) and that community composition is dependent on stochastic processes (i.e., dispersal limitation and ecological drift) causing species abundances to vary. On the other side of the spectrum is niche theory, where species interactions (biotic) and environmental filtering (abiotic) drive community assembly and composition. For niche theory, Hutchinson (1951) and Tilman (1982) provide relatively nice explanations for species coexistence that can be summarized as species can coexist if they can partition their niche along some type of an environmental resource. In particular, the niche-based framework can be a POWERFUL approach because it can account for the links between evolution, (a)biotic environments, and the community [2]. This is because ecological niches are intricately linked to functional traits, as traits underlie an organism's response to abiotic and biotic conditions. Thus, we can examine the evolutionary history of functional traits to gain insights into niche diversification. Recently, these ideas have been explored in microbial ecology. For instance, my dissertation addressed how bacterial ecotypes have differential distributions due to variation in their functional traits [3]. More widely applicable reviews are also available exploring the phylogenetic conservatism of microbial traits [4]. However, we really are at the forefront of understanding the distribution of traits and how evolutionary processes structure microbial trait variability, especially among closely-related taxa. This is why I am really excited to discuss a recent publication concerning the "Emergence of trait variability through the lens of nitrogen assimilation in Procholorococcus" [5]. ProchlorococcusProchlorococcus (Pro) is a globally-distributed marine cyanobacterium that is a major contributor to global photosynthesis. Synechococcus (Syn) is a sister genera that is more abundant at higher latitudes, suggesting partitioning of environmental resources along temperature and nutrient gradients [6]. Within Pro, numerous work over decades in the Chisholm lab have uncovered fine-scale phylogenetic clades that correspond to physiological traits, such as pigmentation, growth rate, and nutrient utilization, and have subsequently been coined ecotypes. By sampling at various depths and across latitudes, Pro ecotypes exhibited differential geographic distributions that were highly correlated to environmental gradients, such as temperature and nitrate [7]. These distributions were corroborated by phenotypic trait assays (i.e., strain that grows better under higher temperatures were found in hotter geographic regions). Thus, by examining trait variation we can better understand the biogeographic distribution of microbes and its relation to niche partitioning. Patchy distribution of Nitrogen traits in ProBoth Pro and Syn are non-N fixing bacterium, and as N can be a limited nutrient for all phytoplankton, it represents a crucial environmental resource. All Syn can assimilate nitrate, but the genetic repertoire in Pro is only found in a few strains that are constrained to a couple ecotypes. This begs the question of whether genomic cluster containing the nitrate assimilation pathway was vertically inherited or if horizontal gene transfer (HGT) mediated the evolution of this trait.
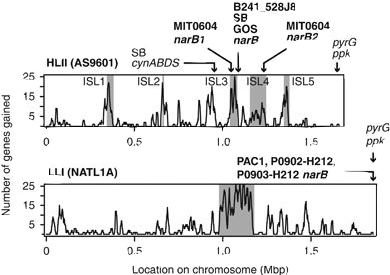
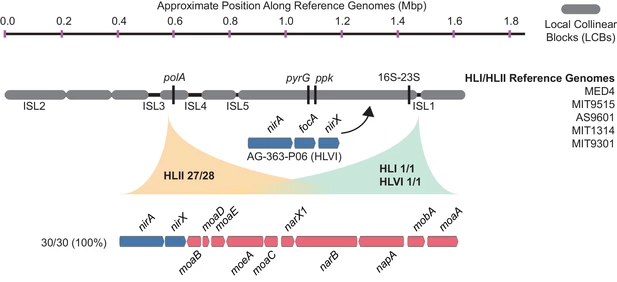
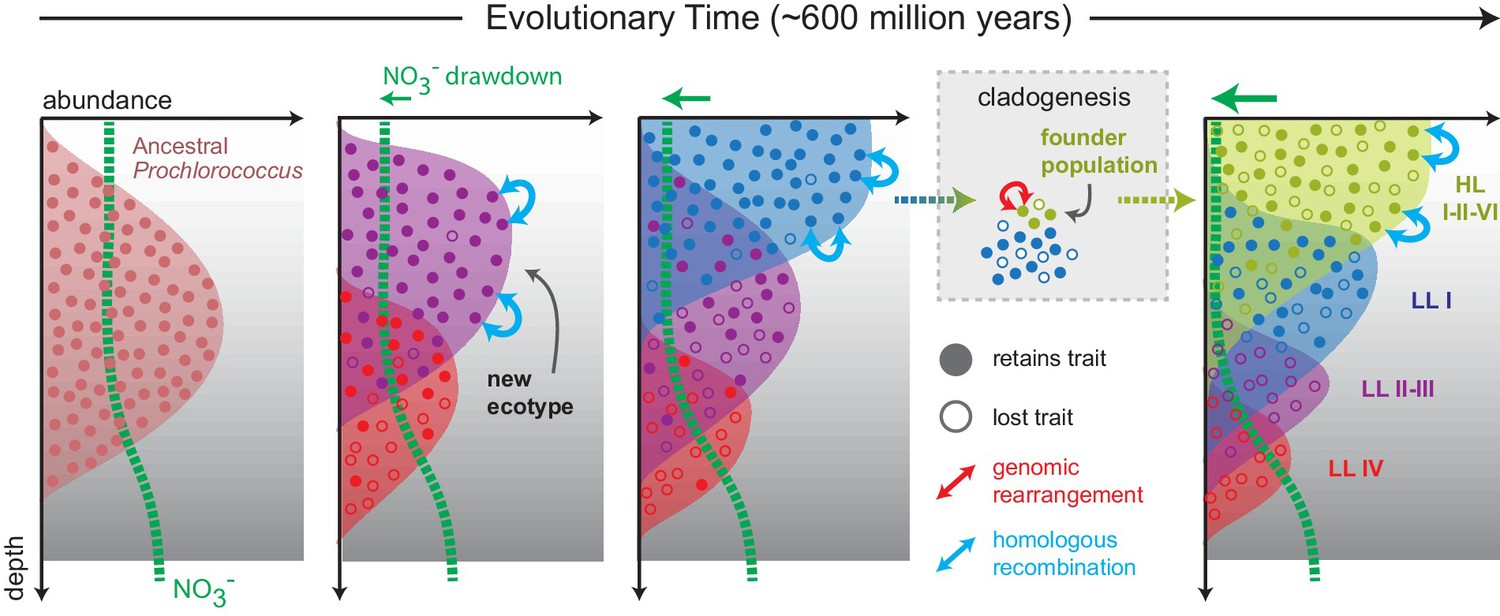
To test this, the authors examined phylogenies for the upstream and downstream genes of the nitrate pathway. Almost universally, the phylogenies were highly congruent with the core genome phylogeny for each gene, rejecting evidence for high degrees of HGT. As further support, comparative genomics of the complete nitrate gene cluster across all Pro genomes revealed high synteny within clades, strengthening the argument for vertical inheritance. Location of the nitrate assimilation cluster in the high light clade (HL). All strains in the HL clade contain identical gene order and genomic location of the nitrate assimilation gene cluster. Homologous recombination shapes trait diversityTo recap, the patchy distribution of the nitrate assimilation gene cluster is not likely due to HGT. Another possibility is the cluster represents a defining trait within Pro clades and provides a differentiating trait within Pro. One such mechanism that reinforces this process is homologous recombination, as the rate of recombination is expected to exponentially decrease with increasing sequence divergence. When the authors investigated the relative role of homologous recombination in structuring the genetic diversity of Pro, they found it can represent a cohesive force shaping the genetic similarity within clades. Indeed, the high r/m (recombination to mutation ratio) suggests that recombination structures genetic diversity far more than mutation accumulation if the genes were diverging. Further, low nucleotide diversity of nitrate assimilation alleles suggests that gene-specific sweeps occurred most likely providing an advantageous trait delineating Pro clades. Evolution of N trait variabilityThe nitrate assimilation gene cluster has a complex evolutionary history that has been mediated through vertical inheritance and high rates of homologous recombination within Pro clades, driving stochastic gene gain/loss to lead to differentiation. However, one major anomaly exists: the absence of the nitrate gene cluster in basal Pro lineages. To address this, the authors finish with an evolutionary model describing the evolution of N and its relation to niche partitioning among Pro clades (see figure below). Briefly, competition and resource trade-offs facilitate partitioning of environmental resources (i.e., nitrate) that causes the stochastic loss of the nitrate gene cluster. When advantageous, homologous recombination has constrained divergence of the nitrate gene cluster even manifesting in gene-specific sweeps within clades. As clades further divergence, homologous recombination is depressed allowing for ecological differentiation. I just want to end with a quote from the paper to illustrate how difficult disentangling these evolutionary processes are; however, as I hope you can take away from this paper, how crucial insights into trait variability can elucidate both ecological and evolutionary processes. "Superficially, this emergent pattern in microbial trait variability might appear to be the result of horizontal gene transfer, but our evidence indicates that the observed patterns can be attributed to processes of vertical descent, gene loss, and recombination between close relatives that have operated throughout the entire radiation of Prochlorococcus." Papers1. Hubbell SP. (2001). The Unified Neutral Theory of Biodiversity and Biogeography. Princeton University Press.
2. Chase JM and Leibold MA. (2003). Ecological Niches. The University of Chicago Press. 3. Chase AB, Gomez-Lunar Z, Lopez AE, Li J, Allison SD, Martiny AC, Martiny JBHM. (2018). Emergence of soil bacterial ecotypes along a climate gradient. Environmental Microbiology 20: 4112–4126. 4. Martiny JBHM, Jones SE, Lennon JT, Martiny AC. (2015). Microbiomes in light of traits: a phylogenetic perspective. Science 350: aac9323. 5. Berube PM, Rasmussen A, Braakman R, Stepanauskas R, Chisholm SW. (2019). Emergence of trait variability through the lens of nitrogen assimilation in Prochlorococcus. eLife 8: e41043. 6. Flombaum P, Gallegos JL, Gordillo RA, Rincón J, Zabala LL, Jiao N, Karl DM, Li WKW, Lomas MW, Veneziano D, Vera CS, Vrugt JA, Martiny AC. (2013). Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. Proceedings to the National Academy of Sciences 110: 9924-9829. 7. Johnson ZI, Zinser ER, Coe A, McNulty NP, Woodward EMS, Chisholm SW. (2006). Niche partitioning among Prochlorococcus ecotypes along ocean-scale environmental gradients. Science 311: 1737-1740. 8. Berube PM, Biller SJ, Kent AG, Berta-Thompson JW, Roggensack SE, Roache-Johnson KH, Ackerman M, Moore LR, Meisel JD, Sher D, Thompson LR, Campbell L, Martiny AC, Chisholm SW. (2015). Physiology and evolution of nitrate acquisition in Prochlorococcus. The ISME Journal 9: 1195-1207.
0 Comments
Leave a Reply. |
AuthorSome thoughts on some (small) things Archives
May 2023
Categories |
Proudly powered by Weebly
 RSS Feed
RSS Feed